Research Topics:
Targeting Burkholderia cenocepacia quorum sensing virulence to fight cystic fibrosis
Burkholderia cenocepacia is a rare opportunistic pathogen, considered a major concern among respiratory tract infections in cystic fibrosis patients. This pathogen is particularly difficult to treat because of its high level of resistance to the clinically relevant antimicrobial agents. In B. cenocepacia, the quorum sensing cell-cell communication system is involved in many processes critical for bacterial virulence, such as biofilm formation, or protease and siderophore production. Targeting the enzymes involved in this process represents a promising therapeutic approach. This research is performed in collaboration with the Molecular Microbiology Group of our Department, led by Prof. Giovanna Riccardi.
|
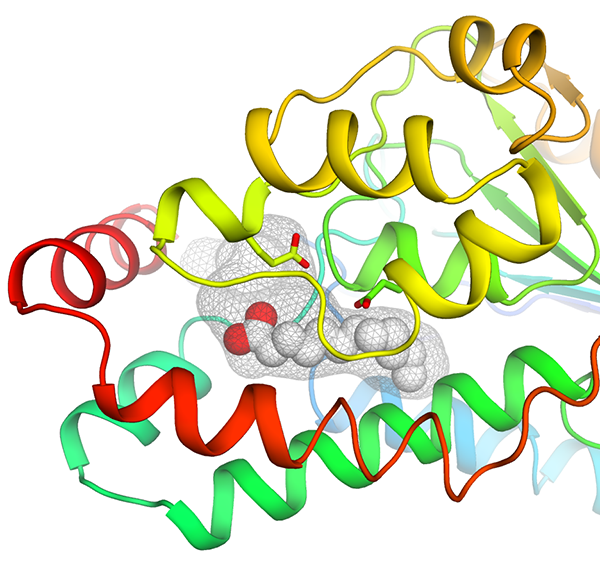
DfsA - The bifunctional crotonase DfsA is a key player in B. cenocepacia quorum sensing pathway. In this work, we presented the three-dimensional structure of the enzyme, fortuitously found in complex with the non- physiological reaction product lauric acid. The biochemical characterization of the enzyme revealed that, differently from the physiological product BDSF ( Burkholderia Diffusible Signaling Factor, cis-2-dodecenoic acid ), lauric acid is slowly released behaving as an inhibitory end product. Collectively, our data illustrate how this enzyme is capable to accommodate long substrate aliphatic side chains, such as its physiological substrate BDSF, without the need for previously suggested conformational changes. These results provide valuable structural templates for future structure-based drug development to target Burkholderia virulence, a trending approach in the fight against bacterial infections affecting cystic fibrosis patients. The figure shows a cartoon representation of the DfsA three- dimensional structure, colored from N-terminus (blue) to C-terminus (red), highlighting the deep hydrophobic cavity (grey mesh). The non-physiological reaction product lauric acid, identified near the catalytic pocket of the enzyme, is shown as spheres. |
|
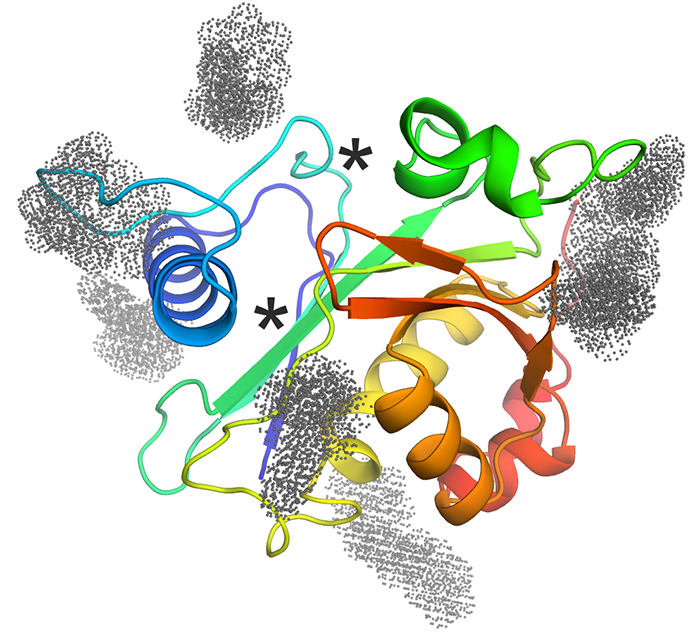
CepI - Our collaborators identified effective non-competitive inhibitors of B. cenocepacia acyl homoserine lactone synthase CepI. Unfortunately, this enzyme refused to crystallize, neither in absence nor in presence of these newly identified inhibitors. Thus, we performed homology modeling and in silico docking of the candidate inhibitors. Using in vitro mutagenesis and biochemical assays, we have validated the docking results to understand their mechanism of action of these powerful candidate drugs. The figure shows a cartoon representation of the CepI homology model, colored from N-terminus (blue) to C-terminus (red). The known substrate binding sites are shown with asterisks. Optimized inhibitor binding sites identified by molecular docking are shown with colored dots. |
PDB Files:
Files in Repository:
Publications:
Frontiers in Pharmacology, 8, 836. (2018) *Equal Contribution
Biochemistry, 55, 3241-3250. (2016) *Equal Contribution - **Shared Corresponding Authors -
PubMed
Scientific Reports, 6, 32487 (2016) *Equal Contribution -
PubMed
PLoS One, 11, pp. e0167350. (2016) -
PubMed
Completed Grants related to this research:
